Researchers identify a new trigger of cellular self-destruction
03-17-2015
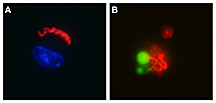
Researchers have identified a bacterial protein that triggers a self-inflicted cell death pathway in immune system cells and could lead to a better understanding of an important cellular structure.
The protein initiates a cascade of events that leads the lysosome, a cellular structure filled with enzymes that break down and digest cellular material, to open holes in its membrane and release enzymes that destroy the cell, said Zhao-Qing Luo, an associate professor of biological sciences at Purdue University who led the research.
"Immune cells have various ways to detect and defend against the presence of pathogens or danger, and cellular death induced by molecules from the pathogens is a common way to fight infection because it cuts off the stream of nutrients and supplies to the pathogen," he said. "However, our understanding of lysosomal cell death is at an early stage despite the fact that the lysosomes play important roles in the development of many diseases. This protein could be a powerful tool to study that process."
The role of the protein, RpsL, was identified in experiments using Legionella pneumophila, the bacteria that causes Legionnaires' disease. The team found that macrophages, a type of white blood cell that recognizes and engulfs foreign entities in the bloodstream, detected the protein when it was released into the cells by the bacetria and initiated cellular suicide to contain the infection. The findings of the study are detailed in a recently published paper in the journal PLOS Pathogens. The study used macrophages from mice, but RpsL is highly conserved among diverse bacteria and appears to be a universal signature recognized by macrophages, Luo said.
The lysosome used to be considered the trash can of the cell, where cellular debris and molecules that had served their roles were sent to be broken down, but it is emerging as an important structure, Luo said.
"It turns out that the lysosome - once considered a waste bin - is actually a treasure box," Luo said. "Abnormal function of the lysosomes has been seen in diseases including cancer and Alzheimer's disease. It is clear we do not know all of the roles it plays and processes in which it is involved, and it could be a valuable target for therapeutics to improve human health."
The lysosome is a sac filled with eznymes that work like knives to slice molecules into smaller subunits that can be reused in the cell. It contains more than 60 different enzymes and is able to selectively release enzymes to accomplish different tasks. How it creates small holes in its membrane and controls which enzymes are released under different conditions is not known, Luo said.
"No one knows how the selective release of the enzymes within the lysosome occurs and no one had been able to induce the process," Luo said. "Now we can. We know that in the presence of this protein, holes open in the lysosome and enzymes are released. This is a critical ability needed to study lysosome function."
The team next plans to try to find the macrophage receptor to this protein and uncover the full cascade of steps and mechanisms involved in the process. The team also will investigate whether the protein triggers the same response in human macrophage cells.
In addition to Luo, co-authors of the paper include Wenhan Zhu and Lili Tao, graduate students in Luo's laboratory at the time of the research; Marsha L. Quick and Johanna A. Joyce from Memorial Sloan Kettering Cancer Center in New York; and Jie-Ming Qu from Shanghai Medical College and Fudan University, in Shanghai, China. The National Institutes of Health and the American Cancer Society funded the research.
Writer: Elizabeth K. Gardner, 765-494-2081, ekgardner@purdue.edu
Source: Zhao-Qing Luo, 765-, luoz@purdue.edu
Original article published in Purdue News March 16, 2015.
Related news releases:
- Purdue scientists reveal how bacteria build homes inside healthy cells
- Purdue biologists identify new strategy used by bacteria during infection
ABSTRACT
Sensing cytosolic RpsL by macrophages induces lysosomal cell death and termination of bacterial infection
Wenhan Zhu, Lili Tao, Marsha L. Quick, Johanna A. Joyce, Jie-Ming Qu and Zhao-Qing Luo
The intracellular bacterial pathogen Legionella pneumophila provokes strong host responses and has proven to be a valuable model for the discovery of novel immunosurveillance pathways. Our previous work revealed that an environmental isolate of L. pneumophila induces a noncanonical form of cell death, leading to restriction of bacterial replication in primary mouse macrophages. Here we show that such restriction also occurs in infections with wild type clinical isolates. Importantly, we found that a lysine to arginine mutation at residue 88 (K88R) in the ribosome protein RpsL that not only confers bacterial resistance to streptomycin, but more importantly, severely attenuated the induction of host cell death and enabled L. pneumophila to replicate in primary mouse macrophages. Although conferring similar resistance to streptomycin, a K43N mutation in RpsL does not allow productive intracellular bacterial replication. Further analysis indicated that RpsL is capable of effectively inducing macrophage death via a pathway involved in lysosomal membrane permeabilization; the K88R mutant elicits similar responses but is less potent. Moreover, cathepsin B, a lysosomal protease that causes cell death after being released into the cytosol upon the loss of membrane integrity, is required for efficient RpsL-induced macrophage death. Furthermore, despite the critical role of cathepsin B in delaying RpsL-induced cell death, macrophages lacking cathepsin B do not support productive intracellular replication of L. pneumophila harboring wild type RpsL. This suggests the involvement of other yet unidentified components in the restriction of bacterial replication. Our results identified RpsL as a regulator in the interactions between bacteria such as L. pneumophila and primary mouse macrophages by triggering unique cellular pathways that restrict intracellular bacterial replication.