|
Research Interests
Structure-Function of the Cytochrome b6f Membrane Lipoprotein Complex
Although there are more than 100,000 independent structures of soluble proteins, and more than 700 such structures of integral membrane proteins (IMP) in the Protein Data Bank, there are only a few structures of hetero-oligomeric oligomeric structures that have been solved to a resolution ≤ 3.0 Å. Although these membrane proteins are of great interest in health and disease-related studies, most of these hetero-oligomeric structures are protein complexes that function in electron transfer in energy transducing membranes. A high resolution (2.7 Å) structure has been obtained of one of the three major hetero-oligomeric membrane protein complexes, the cytochrome b6f complex (see figure below), the electron and proton transferring complex that connects the two reaction center complexes in photosynthetic electron transport.
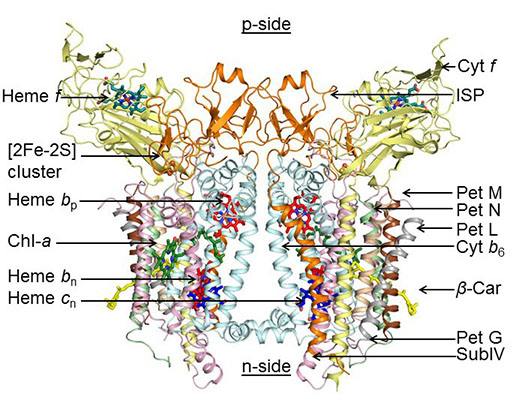
The dimeric 220 kDa complex contains 8 polypeptide subunits (Science, 302, 1009-, 2003; Proc. Natl. Acad. Sci., 103, 69-, 2006; J. Mol. Biol., 370:39-, 2007; J. Biol. Chem., 284:9861-, 2009; Proc. Natl. Acad. Sci., 110,4297-, 2013.) 13 trans-membrane helices, and 7 prosthetic groups (4 hemes, 1 FeS cluster, 1 Chl a, 1 beta-carotene) per monomer. Several structure and function aspects of this complex are novel, including the single chlorophyll a with 2 waters as a ligand, and a covalently bound heme cn, which is 5-coordinate with a hydroxyl ligand bridging to a heme bn propionate, and which is unique in biochemistry because there is no amino acid side chain that serves as an axial ligand. Crystal structures with quinone analogue inhibitors show that heme cn is the electron donor to plastoquinone on the n- or stromal side of the membrane (Proc. Natl. Acad. Sci., 110, 4297-, 2013). The latter manuscript describes the trans-membrane pathway for the H+ transfer that is responsible for most of the photosynthetic trans-membrane proton electronchemical gradient. Among the differences between b6f and respiratory bc1 complexes that imply the mechanism of quinone-mediated electron transfer (“Q cycle”) varies between the complexes is heme cn and the donation of electrons into the b6f complex by the cyclic electron transport chain The details of the structure are of interest for the insight they provide to electron transfer across the dimeric complex, but also for their description of lipid-protein interactions and the labyrinthine pathways of the hydrophobic plastoquinone and –quinol in the cytochrome bc, b6f and bc1 complexes.
Protein Import: The Cytotoxic Colicins
Hypotheses for the pathway and mechanisms of import into E. coli of the cytotoxic E colicins are largely based on structures of a complex of the colicin with its outer membrane vitamin B12 receptor (BtuB), which the colicins parasitize for import. A 2.75 Å structure of the complex of the receptor-binding domain of the endoribonucleolytic colicin E3 showed the elongate 100 Å long colicin domain to be bound in an oblique mode, in which it appears to be 'fishing,' for a second outer membrane receptor-translocator (Nature Struct. Biol., 10, 948-, 2003). A similar structure has been obtained with the receptor binding domain of colicin E2 (see figure) (Sharma et al., JBC, 2007).
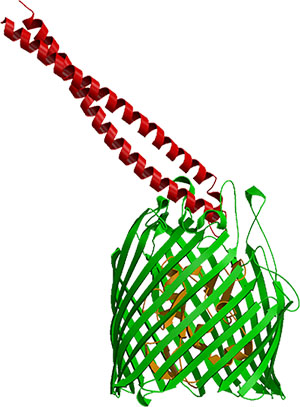
Circular dichroism and electrophysiological studies imply that the colicin is first bound tightly to the extracellular surface of BtuB, is unfolded upon binding, and then uses another very abundant outer membrane protein, OmpF, for translocation (Biochemistry, 45:10199-10207, 2006; EMBO J., 27, 2171, 2008). It is shown that colicin import across the outer membrane involves formation of a ‘translocon’ between BtuB and OmpF. A 3.0 A crystal structure was obtained of a complex between the translocation domain of colicin E3 and OmpF, and 1.6 Å OmpF alone (EMBO J., 27, 2171, 2008). A structure of the BtuB vitamin B12 receptor was obtained, in collaboration with M. Caffrey, by crystallization in the lipid cubic phase (Cherezov et al., 2006). At 1.95 Å resolution, it is the highest resolution structure of a membrane protein not related to bacteriorhodopsin to be obtained by the in meso approach to crystallization.
Lab in 2017: Dr. S. K. Singh, J. Ness, S. Bhaduri, Prof. W. A. Cramer, S. Naurin, S. Colorado Hernandez, Dr. S. D. Zakharov
From the Archives
Moving toward a conceptual break-through
After the canoe trip
Paul Furbacher and I urge Yan-Liang Zhang to study "Forbidden American English"
Sharon Schendel in the lab with rotation student.